Biophotonics............................................................................................................
![]()
Biomolecular analysis methods are foundational to advancing personalized and preventative medicine, accelerating disease diagnostics, and monitoring the health of organisms and ecosystems. High quality factor (High Q) metasurfaces developed in the Dionne group offer a promising platform for rapid optical-based detection and characterization of critical biorecognition elements in health and environment. Metasurfaces are compact arrays of nanostructured elements that, through the design (either forward or inverse) of the scattering components, control the wavefront of electromagnetic fields.
Our High-Q nanoantennas exhibit substantial electromagnetic field enhancements localized around the antennas. The luxury to probe these enhancements through normal free-space illuminations and room for dense biosensor integrations makes this platform viable for building next-generation on-site multiplexed sensors. To know more about the design and fabrication of these High-Q metasurfaces and nanoantennas check Nanophotonics!.
The importance and ubiquity of genetic screening cannot be understated in today’s society. From identification of pathogen nucleic acids like COVID-19 to detection of cancerous mutations to assist in therapeutic treatments, genetic analysis enables significant advances in the prediction, detection, treatment, and monitoring of organisms and health.
Current state of the art technologies such as polymerase chain reaction (PCR), next-generation sequencing (NGS), and Sanger sequencing rely on “traditional” fluorescence or absorbance methodologies, which necessitates time-consuming and expensive sample amplification and replication procedures. Despite their tremendous utility in laboratory settings, translation of such screening methods to clinical and point-of-care applications is impractical.
In our lab, we have designed nanophotonic devices that greatly amplify the light-biomolecule interaction, eliminating the need for traditional sample amplification. By utilizing high-Q metasurfaces with strongly confined modes in small, accessible regions, when decorated with molecular probes through self-assembled surface chemistry, target analyte binding alters the resonator optical signal due to subtle changes in the polarizability or refractive index of the resonator environment.
Furthermore, with the spatially localized nature of our resonances and the choice of our platform (silicon), we can pattern such sensors with high density to allow for analyte parallelizability across a multitude of biomarkers.
While genomics and nucleic acid based diagnostics are incredibly crucial for disease detection and monitoring, they only paint part of the picture of human and population health. To truly advance personalized medicine, our lab is working on utilizing our nanophotonic metasurfaces for protein detection to advance the field of proteomics. There are between 10,000 - 20,000 unique proteins in the human body and being able to rapidly detect thousands of proteins from a single sample is vital for creating an accurate snapshot of the dynamic state of a cell or sample will provide vast amounts of information necessary to truly personalize healthcare. State of the art protein analysis tools and assays include ELISA and mass spectrometry but are limited by their limits of detection and ability to rapidly multiplex samples requiring high-tech equipment, trained personnel, and expensive reagents. Furthermore, unlike nucleic acids, we currently cannot amplify or copy proteins of interest, limiting our ability to detect low concentrations when using traditional optical detection methods such as fluorescence and colorimetrics. As such, our group is working to utilize our high-Q refractive index sensors with specific surface chemistries and acoustic picoliter bioprinting to demonstrate low LoD, multiplexed protein detection.
Photosynthetic microscopic organisms, called phytoplankton, are central to the carbon cycle but under certain conditions can produce powerful bio toxins harmful to both humans and wildlife. In this project, we are developing a high-Q metasurface sensor capable of quantitatively detecting phytoplankton toxins. We fabricate our metasurfaces using Stanford’s state-of-the-art shared facilities, develop chemical assays to specifically bind target analytes to our sensor surface, with an eye towards both selectivity, and specificity, and make our measurements using home-built microscopes that let us test new detection paradigms. In collaboration with the Monterey Bay Aquarium Research Institute and NOAA, we are designing our setup for in-line installation with the Environmental Sample Processor (ESP), an automatic water sampler that rides onboard autonomous underwater vehicles (AUVs) capable of in situ measurements in the open ocean.
Quantitative measurements of biodiversity remain a major challenge. Environmental DNA (eDNA) – the genetic material released from organisms into the environment – is a potential route to detection of harmful, invasive, or endangered species. Current approaches for measuring eDNA such as polymerase chain reaction (PCR) and next-generation sequencing require large capital investment, complex sample processing, amplification of select genes to target organismal groups, and molecular labeling, which can take weeks to months to gain results. We are developing high-Q nanophotonics to quantitatively measure eDNA in marine environments in collaboration with the Micheli Group (Stanford Center for Ocean Solutions & the Hopkins Marine Station) and the Palau National Marine Sanctuary.
Rapid and accurate identification of bacterial infection and antibiotic resistance is essential to improve patient prognosis, slow the spread of infectious diseases, and mitigate antibiotics misuse. Current clinical methods of diagnosis are slow and costly, taking anywhere from hours to days. Bacterial identification based on surface-enhanced Raman scattering (SERS) spectra has been proposed as a rapid and low-cost diagnostic alternative. However, achieving high throughput and attaining reproducible data remain challenging. Additionally, with highly complex SERS spectra originating from a large variety of biomolecules, robust sample processing and data analysis is required for accurate classification. To address these challenges, we are developing colloidal nanoparticle and dielectric metasurface SERS substrates for single-cell measurements at high throughput, chemical modifications of bacteria to increase selectivity of SERS enhancements, and acoustic bioprinting for picoliter sample separation. Our research is bolstered through interdepartmental collaborations with the School of Medicine, Graduate School of Business, and Departments of Computer Science, Electrical Engineering, and Mechanical Engineering.
Different species of bacteria have unique Raman “fingerprints” describing the chemical composition of their cell surfaces. This variation holds enormous potential for rapid diagnosis of bacterial infections using Raman spectroscopy. We have collected Raman spectra for 30 species responsible for a majority of the world’s bacterial infections and demonstrated that deep learning can classify bacterial species based on their Raman fingerprints with more than 80% accuracy. To further improve this system, we are using SERS to overcome the intrinsically low efficiency of Raman scattering. We are using gold nanorods as our SERS substrate, which can be tuned to resonate with the incident laser, and selectively enhance bacterial signal by electrostatically binding to them. With surface enhancement, Raman fingerprints are clearly discernible with less than 10 second exposure, enabling bacterial identification in a matter of seconds.
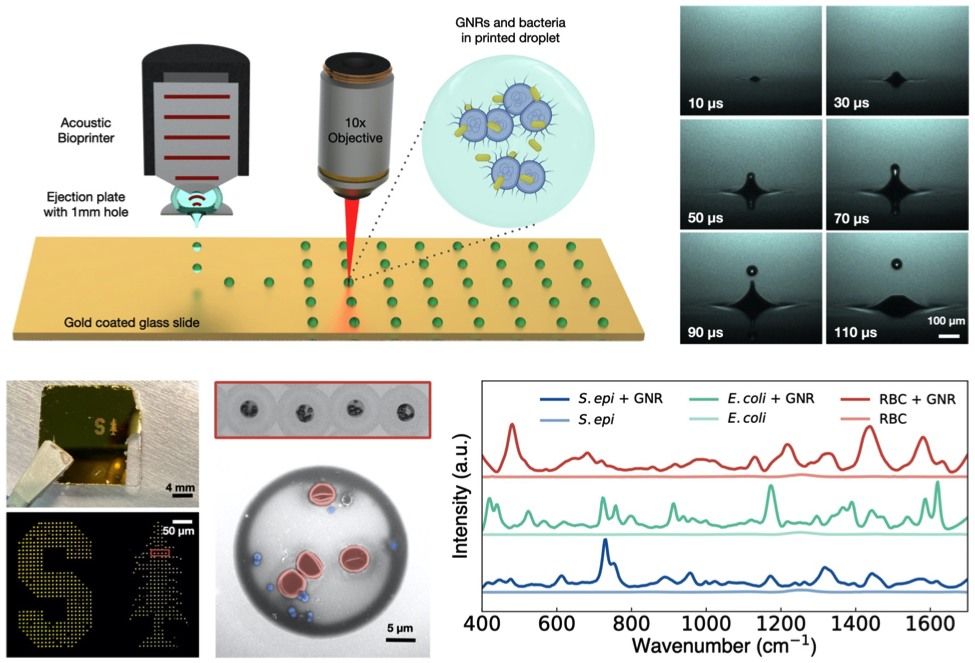
Surface-enhanced Raman spectroscopy (SERS) is a promising approach for sensitive, specific, and rapid culture-free cellular identification. While promising in its specificity, SERS-based platforms suffer from challenges in sample preparation. In our lab, we demonstrate the use of an acoustic droplet ejection platform that generates droplets containing cells uniformly coated with gold nanorods (GNRs) while achieving minimal GNR loss. We have shown that we can stably eject ~2pL droplets from diverse solutions containing S. epidermidis, E. coli, and mouse red blood cells mixed with GNRs at 1 kHz ejection rates. From these samples, we can collect Raman spectra within 15 seconds and demonstrate spectral enhancements up to 1500x as compared to droplets printed without GNRs. Additionally, we employ machine learning algorithms to classify our spectra. Using a Random Forest classifier, we have demonstrated 99% classification accuracy of droplets printed from single cell lines and >82% classification accuracy of droplets printed from mixtures of RBCs and bacteria. As such, we’re excited about the potential for combining acoustic droplet ejection with SERS and machine learning to push forward the clinical and industrial translation of SERS-based cellular identification.